QC L2 criteria¶
- Table of contents
- QC L2 criteria
General¶
Three levels of data granularity are involved in the quality checks of level 2 (QC L2) and the assignment of QC level 2. These are in ascending size:- Atomic Dataset: This is the granularity to which the exceptions thrown by the qc2 tool refer. The levels of exceptions are as listed and categorised in file#3246 .
- ESG Dataset: The accessible data unit in the ESG gateways (consisting of atomic datasets).
- Experiment (consisting of ESG datasets): At this data granularity the quality levels are assigned. This is also the data granularity of the DOI assignment.
The presented QC L2 assignment criteria are general rules but no simple check list, since automated QC checks can only give hints for problems in the data, but a scientist has to judge them. E.g., constant value fields for wind components or temperature distribution are erroneous but for snow heights it might be sensible.
QC L2 exceptions: severity levels and thematic groups¶
Groups of severity levels¶
The exceptions were grouped by severity into five severity levels.See file#3235 for an exception list ordered by exception keys. There the colours indicate the different severity levels.
- Fatals: immediate action neccessary,
- errors: data not acceptable,
- warnings: data possibly not ok,
- informatory: fine - just for info,
- unclear: title lines or open issues.
Thematic groups of exceptions¶
The exceptions were grouped into seven thematic groups, as well. This grouping is for practical use and overview only. It presently has no effect on workflow or other behaviours. These groups are (abbreviations in capitals):- ACCEss errors,
- GENEral checks,
- METAdata and file name checks,
- TABL: inconsistencies in comparison to meta data tables (CMIP5 standard table or project table),
- TIMEaxis checks,
- VARIables' checks,
- obsolete messages (e.g., not used for CMIP5 project).
See file#3235 for an exception list ordered by thematic groups.
QC L2 assignment procedure¶
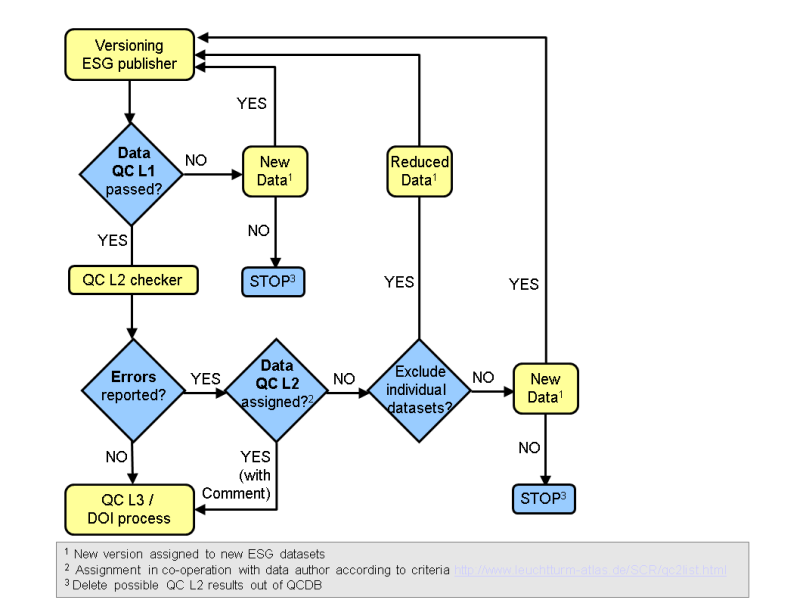
Analysis of exceptions for atomic datasets¶
- Get the exception statistic of your QC run [analyse log files of qc tool]:
qcDbselect.py --postgresdb=<qcdb> --experiment=<drs experiment> [--logout=<download dir>]
- qcDbselect_<drs_experiment>.log: list of atomic datasets with detailed exception message (also stdout) and count of highest exception categories per atomic dataset
- QCL2_statistics_<drs_experiment>.txt: list of all exceptions and the atomic datasets, where they occurred
Note: There might be multiple different (or same) exceptions detected for a single atomic dataset. Thus the total number of entries of this list is higher than the number of checked atomic datasets.
- Atomic datasets with one or more exceptions classified as "FATAL" need interaction as
- the qc tool might have stopped the checks of this atomic dataset (e.g., due to lack of access rights),
- the exception might have occurred due to improper usage of the qc tool (e.g., due to erroneous entries in the setup file),
- the data might be corrupt (e.g., due to severe errors). In this case the data should be rejected.
- For exceptions of the category "ERROR" the data need inspection and the data author should be contacted. Data may be resent.
- About exceptions of the category "WARNING" the data author should be informed. Normally, they are just informative, but they might provide some hints for other exceptions detected.
- The data author has to be involved for any suspicious data. If the data author can explain exceptions as normal behaviour of his/her model, the data can still be accepted. This has to be documented during QC L2 assignment (see below).
ESG Dataset level¶
The QC results for the atomic datasets are aggregated on ESG dataset level. Three different cases are to be distinguished:
- PASSED: In case no unacceptable exceptions have occurred in any of the atomic datasets belonging to an ESG dataset, this ESG dataset has passed the QC L2 checks.
- FAILED: In case of unacceptable exceptions in all atomic datasets, the ESG dataset has failed the QC L2 checks. The author has to be informed. It has to be discussed with him, whether these data should be delivered again or this ESG dataset is to be excluded from the QC procedure.
- In case some but not all of the atomic datasets failed the QC L2 checks, we suggest that in agreement with the data author
- either the faulty parts of the data will be replaced (restart of overall QC procedure afterwards)
- or these atomic datasets are excluded from QC process, i.e., the remaining atomic datasets are ESG published under a new version. The QC tool is applied to this version (QCDB_VERSION=<assigned version> ). For safety reason deletion of QC results for excluded data (using qcDbdelete.py). The reduced ESG dataset then passes QC L2 checks.
QC L2 assignment level: experiment¶
We can distinguish three similar cases in the aggregation of QC results on ESG dataset level to experiment level than from atomic dataset level to ESG dataset level:
- PASSED: In case all ESG datasets belonging to the experiment have passed the QC L2 checks, the experiment has passed QC L2, too, and can be assigned.
- FAILED: In case all ESG datasets failed the QC L2 checks, the experiment has failed QC L2 checks. There is no QC level 2 assigned to the experiment, i.e., data is rejected. The author has to be informed. It has to be discussed with him, whether these data should be delivered again or is to be excluded from the QC procedure (i.e. no DOI assignment on this experiment).
- In case some but not all of the ESG datasets failed the QC L2 checks, we suggest that in agreement with the data author
- either that a part of his data will be replaced (restart of QCL2 procedure)
- or that the failing ESG datasets are excluded from QC procedures (no new qc tool run necessary) and the reduced experiment data volume is assigned QC level 2 (passed).
- Exclusions of data have to be done in agreement with the data author.
- The assignment of QC L2 implicitly includes all ESG datasets in the newest available version in the QC database (which is QC L2 checked).
The assignment of QC L2 to an experiment is by:
qcAssignL2.py --postgresdb=<qcdb> --experiment=<drs experiment> --cimxml=<cim quality report xml> --plot=<qc result plot.pdf> --comment=<comment on QC L2 assignment>
Notes:
- Please specify any data exclusions from your QC L2 assignment that were done during the QC assignment procedure using the option 'comment' of the qcAssignL2 command. We need that information for QC level 3 and the DOI assignment.
- In case you have data with thrown errors PASSED, please specify what data and the reason why they were accepted.
- We recommend to provide a qc result plot along with your qc results (option 'plot')